

We are dedicated to providing outstanding customer service and being reachable at all times.

Human Whole Genome Sequencing
Whole genome sequencing (WGS) has focused on the analysis of single nucleotide variants (SNV) and small fragment insertion-deletion mutations (InDel), the detection of variants in some repetitive sequence regions, and the detection of structural variants (SV) in large segments, with far-reaching implications for the pharmaceutical, environmental and biotechnology industries. Since traditional short-read sequencing technology does not typically span entire regions of interest, including repeats and structural variants and full-length RNA transcripts, WGS based on next-generation sequencing technology faces challenges in detecting the above items.
WGS has become a widely used method for studying variation in the human genome. CD Genomics is now providing PacBio SMRT sequencing for human whole genome sequencing. Whole genome sequencing of different human individuals or populations using PacBio SMRT sequencing and perform bioinformatic analysis at the individual or population level. It can comprehensively explore the SNP/ InDel/ copy number variation(CNV)/ SV variants in the genome and provide important information for screening disease causative and susceptibility genes and studying pathogenesis and genetic mechanism.
Project Workflow

Sample Requirements
- DNA amount: ≥ 10μg
- DNA Purity: OD260/280 =1.8 ~2.0 without degradation or RNA contamination
Sequencing system: the PacBio Sequel II or IIe sequencer.
Sequencing Strategy: 20kb Library, ≥ 10X genome coverage depth.
Features of The PacBio Sequel II System

-Automated consumable handling with integrated software.
-Intuitive run setup and monitoring.
-Flexible run time.
-Walk away time up to 4 days.
-Ideal for projects such as rapidly and cost-effectively generating high-quality whole genome de novo assemblies.
Bioinformatics Analysis
The genomes will be assembled using full-solution analytical software tools and standard file formats. Our customized bioinformatics services are also available upon request, generally including data quality control (QC), reference genome alignment, genome annotation, variation annotation, disease genome association analysis, and so on.
Analysis Pipeline
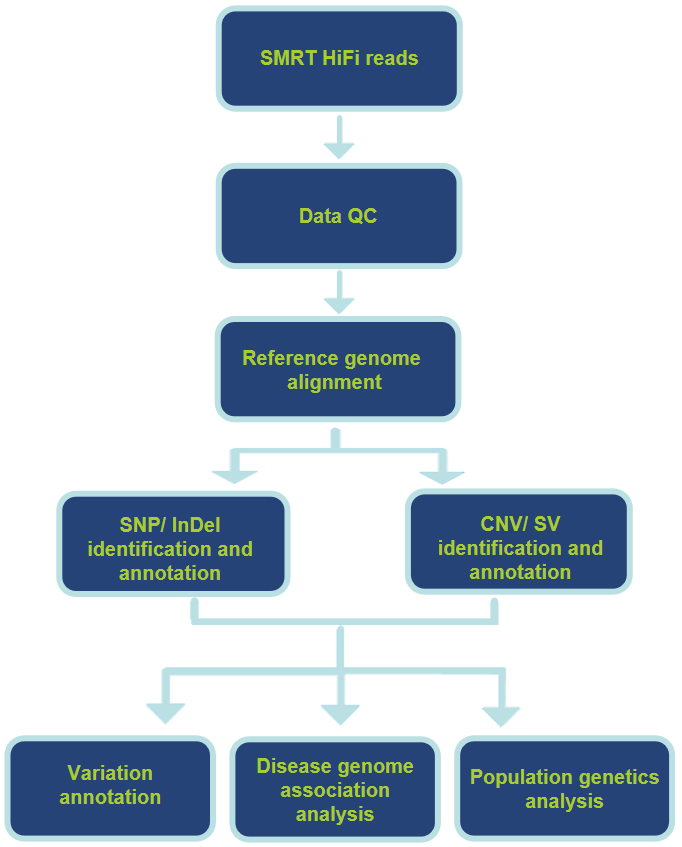
Our Services Support
- To produce reliable references for a population, individual, or diseases (e.g. cancers, monogenic diseases, and complex diseases).
- To identify common alleles in the population by generating fully phased haplotypes. A haplotype, also known as Haploid genotype, is a set of alleles in an organism that is inherited from a single parent.
- Access to hard-to-characterize regions and new types of genetic variation.
- To improve variant detection at population-specific loci using new reference sequences.
Benefits of Our Services
-Exceptionally long read lengths (20 kb or more) and unparalleled consensus accuracy.
-Meet the needs of multiple sample types, such as blood samples, saliva samples, genomic DNA samples, routine cryopreserved fresh tissue samples, etc.
-Decades of experience in genomics services and innovative technology platforms.
-Flexible sequencing strategies and bioinformatics pipelines.
Through SMRT HiFi sequencing, CD Genomics can avoid the problems of high GC regions that cannot be spanned, high repeat fragments that cannot be accurately quantified, and rare mutations that are overwhelmed, and thus generate reference-quality assemblies of diverse populations to better understand the complexity of human health and disease. If you are interested in our services, please contact us for more details.
Reference
- Rhoads, A., & Au, K. F. (2015). “PacBio sequencing and its applications.” Genomics, proteomics & bioinformatics, 13(5), 278-289.
For research purposes only, not intended for personal diagnosis, clinical testing, or health assessment