

We are dedicated to providing outstanding customer service and being reachable at all times.
Human Leukocyte Antigen (HLA) Typing
Human leukocyte antigens (HLA), often referred to as the major histocompatibility complex (MHC), are a group of genes that contain the most polymorphic loci in the human genome. HLA regions are associated with the largest number of human diseases in the human genome, and HLA typing is particularly important for the evaluation of immunologic compatibility between organ donor and recipient pairs. The investigation of the polymorphism of HLA is necessary and challenging due to its highly polymorphic. CD genomics is based on third-generation sequencing technologies, including Oxford Nanopore Technologies (ONT) and Pacific Bio (PacBio), which supports high-resolution HLA typing in a rapid and cost-efficient manner that contributes to a better understanding of HLA function, its causal relationship to disease, and so on.
Overview of Human Leukocyte Antigen (HLA) and Its Typing
HLA is present in all jawed vertebrates, and for humans, it is located in a region approximately 4 M long on the short arm of human chromosome 6 (6p21.3) and has more than 200 protein-coding genes. The HLA gene products can be expressed on different cell surfaces and play critical roles in antigen presentation and immune signaling. HLA is closely related to disease defense, organ transplant rejections, and drug adverse reactions. Therefore, HLA has been widely applied in susceptibility gene detection for immune-related diseases, hematopoietic stem cell transplantation (HSCT), and drug allergy testing. There are mainly three regions of HLA, including HLA class I genes (e.g. HLA-A, HLA-B, and HLA-C), HLA class II genes (e.g. HLA-DP, HLA-DQ, and HLA-DR), and HLA class III genes. Among them, the first two classes play critical roles in immune responses, with HLA class I genes encoding antigens primarily for CD8+ T cells and HLA class II genes encoding antigens primarily for CD4+ T cells.
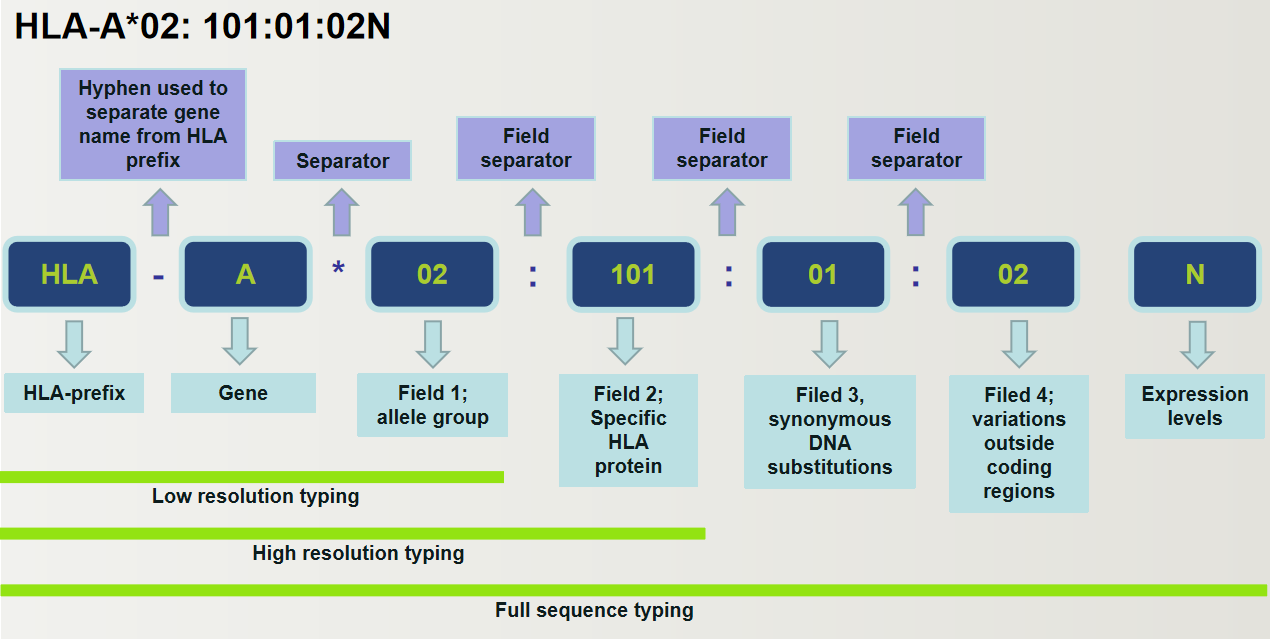
The resolution of HLA typing has a significant impact on HLA matching, and the main categories are as follows:
-Low resolution typing (also known as first field typing): a set of HLA alleles (allele family), equivalent to serological typing.
-High resolution typing (also known as second field typing): refers to one or a group of HLA alleles for the same antigen-binding site.
-Allele level typing (also known as field typing): the exact nucleotide sequence of an HLA gene.
-Other levels of resolution: intermediate levels of typing, where specific subtypes can be defined.
We Provide a Complete End-To-End Solution

Sample Requirements
- Start with high-quality, genomic DNA, as low as 250ng to 1µg.
- Other sample types, including blood, tissue, cell lines, and so on.
Our Analysis Service Workflow
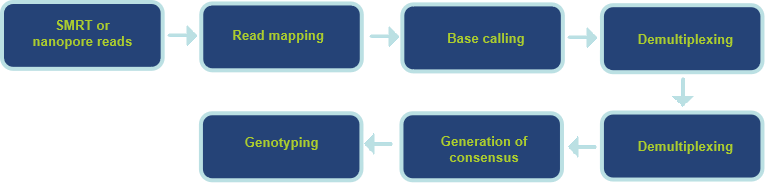
Our analysis services include basic bioinformatics and advanced bioinformatics services, depending on the client's requirements for the project, time, and budget. The basic bioinformatics services provide de novo HLA gene assembly, HLA annotation, and HLA analysis. The advanced bioinformatics services include shared HLA type screening, significance analysis based on statistical tests, and more.
Advantage of Our Services
- The ability to assemble the MHC region and phase the entire MHC region of 3.6 Mb.
- Helping elucidate information on alleles, the localization of mutant loci, and non-coding regions on each MHC haplotype.
- Highly accurate HLA typing to meet the needs of studies such as autoimmune diseases and pharmacogenomics, etc.
- Allowing significantly shorter turnaround times for multiple samples at a lower cost.
Notably, third-generation sequencing (TGS) technologies offer a significant advancement over current next-generation sequencing (NGS) platforms for all HLA genotyping needs. Our services based on nanopore sequencing and SMRT sequencing can deliver HLA typing in a simple, rapid, and cost-effective manner. Please don’t hesitate to contact us. Discuss your projects with our professional scientists.
References
- Liu, C., et al. (2018). "Accurate typing of human leukocyte antigen class I genes by Oxford nanopore sequencing." The Journal of Molecular Diagnostics, 20(4), 428-435.
- Johansson, T., et al. (2021). "Targeted RNA-Based Oxford Nanopore Sequencing for Typing 12 Classical HLA Genes." Frontiers in genetics, 12, 273.
For research purposes only, not intended for personal diagnosis, clinical testing, or health assessment