

We are dedicated to providing outstanding customer service and being reachable at all times.
Animal/Plant Whole Genome De Novo Sequencing

Plant and animal de novo sequencing refers to the sequencing of a new genome by bioinformatics analysis without any reference sequence, which is aligned, spliced, and assembled to generate the initial genome sequence of a specific organism. Through this method, the whole genome sequences of plants and animals can be obtained, and then a series of studies such as functional gene mining, regulatory metabolic network construction, and species evolution analysis can be carried out downstream of the species to advance the research of the species. Based on the features of long read length, no GC preference, and high sequencing accuracy, CD Genomics can help our customers decipher most of the structural properties of the genome using long-read technologies.
Advantages of De Novo Sequencing with Long-Read Technologies
-PacBio SMRT Sequencing
PacBio SMRT Sequencing technology has two sequencing modes, including continuous long read (CLR) and circular consensus sequencing (CCS). Among them, CCS can generate High-fidelity (HiFi) reads, the latest data type developed by PacBio with long reads and high accuracy. As a result, PacBio HiFi reads can greatly enhance the accuracy of genome assembly, even for the most complex genomes.
-Oxford Nanopore Sequencing
Nanopore sequencing differs from other sequencing platforms in that the nucleotides are read directly, eliminating the need for a DNA synthesis process. Moreover, the nanopore sequencing platform itself has no significant technical limitations on DNA read length. Ultra-long read lengths (ULR, N50=100-200Kb, one of the data types generated by nanopore sequencing) can be utilized for high-quality genome assembly.
Our Technology Pipelines
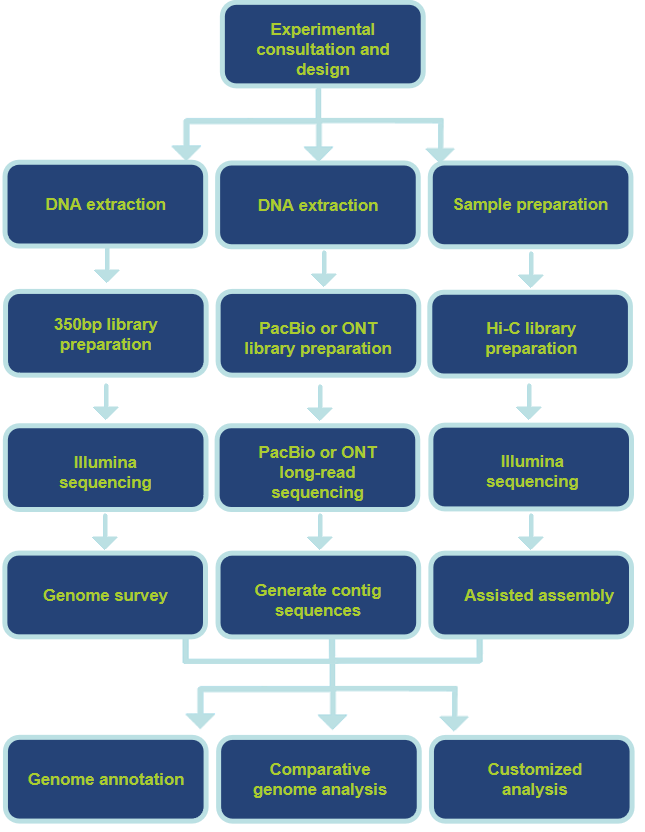
Sample Requirements
- Genomic DNA amount:
Illumina NovaSeq platform: Genomic DNA ≥ 5μg, OD260/280 =1.8 ~2.0 without degradation or RNA contamination.
PacBio Sequel II sequencing platform: HMW Genomic DNA≥ 15μg, OD260/280 =1.8 ~2.0, A260/230=1.5 ~2.6
Nanopore PromethION sequencing platform: HMW Genomic DNA≥ 8μg, OD260/280 =1.75 ~2.0, A260/230=1.4 ~2.6 - For additional sample inquiries, please contact our professional technicians.
Bioinformatics Analysis Contents
| Items | Contents |
| Genome assembly | -Assembly strategy -Assembly result evaluation |
| Genome annotation | -Gene structure prediction and annotation -Gene function prediction and annotation -Repetitive sequence prediction and annotation -Non-coding RNA prediction and annotation -Pseudogene prediction and annotation -Transposon classification |
| Comparative genome analysis | -Gene family clustering -Phylogenetic tree construction -Gene family expansion and contraction analysis -Specific gene family analysis -Whole genome duplication event analysis -Selective pressure analysis |
| Customized analysis | -Customized analysis based on specific client projects |
Application of Animal/Plant Whole Genome De Novo Sequencing
- Building species databases.
- Study of the evolutionary origin of species, trait characteristics.
- Mining functional genes.
- Laying the foundation for subsequent resequencing to discover single nucleotide polymorphisms (SNPs) and other genetic variants.
Long-read technologies combined with other complementary technologies have become the technology of choice for plant and animal genome assembly. CD Genomics' services typically include experimental consultation and design, sample preparation, multi-platform sequencing, read assembly, genome and functional annotation, advanced bioinformatic analysis, and customized analysis for specific biological queries. If you are interested in our services, please don't hesitate to contact us. Discuss your projects with our professional scientists.
For research purposes only, not intended for personal diagnosis, clinical testing, or health assessment